Omics Integrator
Omics Integrator is package comprised of command-line tools designed to integrate high-throughput datasets such as gene expression, phospho-proteomic data and the results from genetic screens. As shown below, Garnet is used to identify transcription factors that give rise to gene expression changes using epigenetic data while Forest integrates these data or other data by finding connections in a protein interaction network.
PIUMet
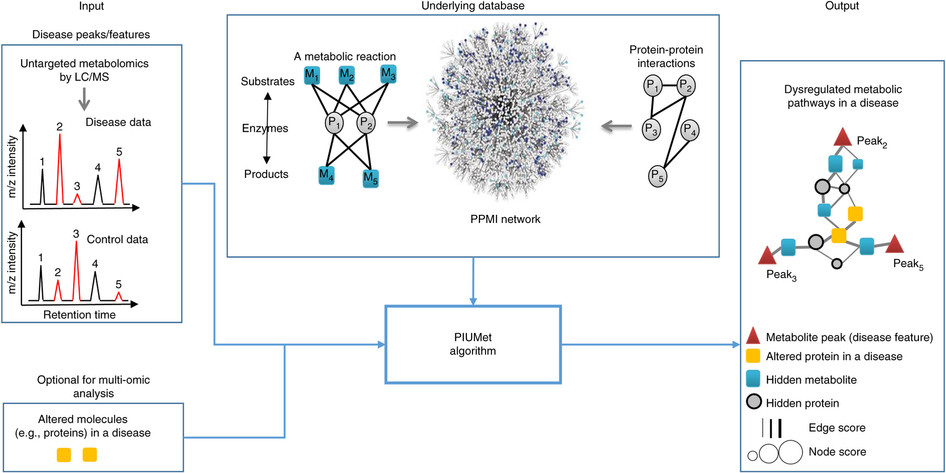
PIUMet is a network-based algorithm for integrative analysis of untargeted metabolomic data. It leverages known metabolic reactions and protein-protein interactions to analyze the ambiguous assignment of metabolomics features and identify disease-associated pathways and hidden components.
NeuroLINCS Galaxy
We’ve released a custom ‘flavor’ of Galaxy, a docker image producing a galaxy optimized for our analyzes, available to the community.
The NeuroLINCS-Galaxy Docker Image. If you intend to reproduce or conduct similar analyses at scale, comes pre-configured with the tools we use as part of our workflows.